Since the launch of the project in March 2019, participants of EOSC-Life have been working hard to get data, tools and workflows from the life sciences ready for deployment in the European Open Science Cloud. Please click on the image of our final Achievements Brochure to read about the project outcomes.

EOSC-Life swiftly responded to the challenges presented by the COVID-19 pandemic, helping partners to develop three critical resources:
In response to the pandemic, EOSC-Life helped partners develop a repository to store Individual Participant Data (IPD) from clinical trials using the TSD platform for sensitive data. Integrated in the European COVID-19 Data Portal, this repository is certified, subject to rigorous governance, ensures long-term data preservation, and complies with relevant regulations (e.g. GDPR), facilitating data re- analyses, secondary analyses, and patient-level meta-analyses.
Extension of the COVID-19 Data Portal
Because >75% of the world’s data flows through the Data Hubs, EOSC-Life extended the COVID-19 Data Portal to mobilise open biomolecular data (500,000+ records from the biomolecular and literature domains) and new SARS-CoV-2 data (>160,000 viral isolates with raw sequence data), connecting these data with clinical and epidemiological data. This, in turn, has facilitated data sharing and analysis and accelerated coronavirus research.
Clinical Research Metadata Repository
EOSC-Life helped develop the Clinical Research Metadata Repository (MDR), including COVID-19 data. Researchers can use the MDR to access clinical trials and related data objects (protocols, info sheets and consent forms, data management plans, statistical analysis plans, results, publications, and descriptive metadata). The MDR makes clinical research data from all disease areas FAIR by increasing data Findability.
View even more resources from EOSC-Life here.

EOSC-Life helped ESFRI Life Science RIs develop repository infrastructure in many ways, building data expertise in LS RIs to expand the community of experts in these infrastructures.
EOSC-Life funded 8 “Demonstrator” projects representing novel cloud deployments from participating RIs . These scientific and technical use cases guided and structured the work done in EOSC-Life, showing how an open digital and collaborative space for biological and medical research can be built. Covering a broad scope of life science domains, these projects made new data, tools, and workflows available in the cloud for re-use by the scientific community. The publications and resources, including training material and documentation, are openly accessible.
EOSC-LIFE created an open, collaborative digital space for the life sciences in the EOSC by making RI data and services available in accordance with the FAIR principles (findable, accessible, interoperable and reusable). Eight (8) pilot projects were supported, representing a wide range of data commonly used by life sciences user groups, to renew these data and services. This renewal helped to ensure the sustainability of these resources and their use.
You can find all videos of the presentations on this page.
Phytoliths, silica bodies deposited in or between plant cells during the plant’s life cycle, are used in archaeology, palaeoecology, and the plant sciences to document changes in environmental and biodiversity. EOSC-Life funding supported the FAIR Phytoliths Project, increasing the FAIRness of phytolith data, methods communication, data sharing, and archiving practices. An open repository for phytolith data was also created, populating the EOSC with novel data in this discipline.
View even more resources from EOSC-Life here.

RO-Crate is a method that was established in EOSC-Life as a community standard to practically achieve the FAIR packaging of research data with their structured metadata. The EOSC-Life project significantly contributed to RO-Crate development, enabling it to support the workflow life cycle. This cycle includes (meta)data deposition and archiving in WorkflowHub (a European-wide repository of computational workflows in of life sciences), a broader spectrum of workflow execution processes (WfExS, SCHeMa), monitoring (Life Monitor), reporting (OpenEBench), provenance (Common Provenance Model), and workflow submissions for the regulatory sciences (BioCompute Objects). EOSC-Life also aligned RO-Crate with FAIR Digital Objects through the Research Data Alliance and the DISSSCo SYNTHESYS+ project.
RO-Crate is supported by open source tooling and is now being adapted for use outside the life sciences, in application areas as diverse as the cultural heritage archives (e.g. PARADISEC) to Machine-actionable Data Management Plans (e.g. maDMP) and institutional repositories based on Dataverse (e.g. Harvard Data Commons). Within the EOSC, RO-Crate is also used by the earth sciences for data cubes (RELIANCE) and for data collaboration (CS3MESH4EOSC). Anyone can join the RO-Crate Community, and members meet regularly to consider new use cases, improve the specifications and tooling, and coordinate implementation actions across EOSC projects.
Go to RO-Crate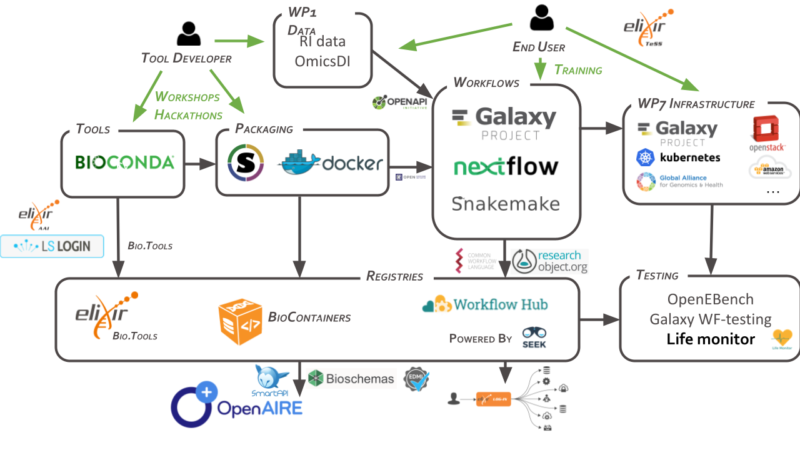
EOSC-Life developed expertise in the cloud deployment of software and workflows across all life science domains. This helped LS RIs develop their computational infrastructures still further and make them FAIR. EOSC-Life first identified technologies and standards that could be readily supported within the project, resulting in the creation of a Tools Collaboratory Roadmap.
In EOSC-Life, this Roadmap promoted interoperability across the LS RIs and domains, while less mature infrastructures could use it to coherently shape their own computational infrastructures.

Learn more about workflows and tools applied and developed in EOSC-Life here.
EOSC-Life developed this registry dedicated to describing, sharing, and publishing scientific computational workflows. The registry makes it easier for tool developers and users to discover and re-use workflows in accessible and interoperable ways through the extensive use of open standards and tools, including CWL, RO-Crate, Bioschemas, and GA4GH’s TRS API, in accordance with the FAIR principles.
Due to the COVID-19 pandemic, EOSC-Life shifted its focus towards providing tools and workflows that could be used to analyse COVID-19 data transparently and reproducibly. The WorkflowHub was accelerated by 6 months and released early to provide a registry for COVID-19 workflows. Thanks to a collaboration with the global Galaxy community, these workflows are now available on public Galaxy instances worldwide.
Go to WorkflowHub
EOSC-Life WP2 developed the LifeMonitor service to support the sustainability and reusability of published computational workflows. Since the first public release in May 2021, the service has continuously progressed in functionality and has been adopted to monitor the workflows of the Galaxy IWC collection. This service also offers a web application that allows users to easily register workflows from WorkflowHub and other sources and to easily track the test status of multiple workflows and apply workflow best practices.

In order to be able to find, use, or interpret data and software in the long term, they need to be archived in online portals with their associated detailed and semantically annotated metadata. EOSC-Life helped to identify specific semantic data interoperability challenges, expand domain coverage, and even develop a unique identification system based on Digital Object Identifiers (DOIs) for specific life science resources. One of EOSC-Life’s primary goals was to support the registration, dissemination, and application of reference collections of curated FAIR data repositories, such as those hosted by FAIRsharing.org.
FAIRsharing.org is a curated, informative, and educational resource on data and metadata standards related to databases and data policies. It also features a prototype educational page on standards. The resource serves all disciplines and has been adopted by funders, publishers, RDA, and other organisations. The EOSC-Life FAIRsharing collection includes more than 130 diverse data resources produced by EOSC-Life partners.
EOSC-Life contributed to several additional sustainable outcomes related to improving the FAIRification of data:
EOSC-Life launched the FAIRassist.org website, placing a strong focus on listing and describing existing resources that could be used for the assessment and/or evaluation of digital objects with reference to the FAIR principles.
EOSC-Life provided support that enabled the development of the FAIR Cookbook, an online, open and live resource for the life sciences containing “recipes” that help users to make and keep data Findable, Accessible, Interoperable and Reusable (FAIR).
EOSC-Life provided support, development, and hosting for the automated FAIR Evaluator.
The Terms4FAIRskills initiative was launched with an EOSC-Life WP9 Training, which instructed users on how to build a terminology for the skills and competencies necessary to make data FAIR and keep it FAIR.
EOSC-Life created a comprehensive document that provides a concise overview of FAIR requirements.
EOSC-Life updated specifications and documentation on the FAIR data point.
These were identified and discussed in a special issue of the Data Intelligence journal.

EOSC-Life published new standards and established new services that protect and support the existence and sustainability provenance information provided via metadata, thus enabling data (object) history to be traced and preserved. EOSC-Life facilitated the development of a Common Provenance Model, introducing a framework to handle distributed provenance information. Throughout the project, the systematic use of RO-Crate and LifeMonitor have supported provenance management.
EOSC-Life supported the development of a Common Provenance Model and published a standard under ISO/TS 23494-1 to describe the history of (sensitive) data in the life sciences in distributed environments. The model thus enables these data to be assessed for their reusability in further research, improves the reproducibility of research results, and builds bridges between institutions. The standard also supports compliance with the Nagoya Protocol
EOSC-Life developed a model that could be used to conduct a privacy risk analysis of whole-slide images, placing a specific focus on how to guard against identity disclosure attacks, as these are the most important from a regulatory perspective. Guidelines were also provided on how to anonymise these data, which had been a long-term unsolved problem in the field of digital pathology.
Access the publication describing the model and guidelines in Nature Communications
Sensitive data generate an additional layer of complexity for life science infrastructures. Interoperability challenges such as cross-domain categorisation and the discovery of digital resources related to sensitive data must be overcome. To support the FAIRification of sensitive data resources, EOSC-Life developed a toolbox demonstrator to make it easier to discover digital objects related to these data and thus to use them.

EOSC-Life implemented a suite of semantic, FAIR interoperability services, helping to ensure that (meta)data use a formal, accessible, shared, and broadly applicable language to represent knowledge.
This repository for biomedical ontologies provides a single point of access to the latest ontology versions. Researchers can browse the ontologies through the website as well as programmatically via the OLS API. These relationship-based hierarchical descriptions within biomedicine capture a common language for reuse and reapplication in computer systems, improving efficiency and flexibility between systems, and aiding in language processing.
The OxO Service provides a database of ontology cross-references (or xrefs) extracted from multiple sources, including ontologies in the OLS. Users can upload their own mappings, search the OxO database for mappings for a given identifier using a range of identifier types, or share mappings created for specific contexts. EOSC-Life implemented the OxO in the form of a suite of semantic services, deployed using cloud principles and technologies.
EOSC-Life developed and harmonised life science community definitions of metadata that are required to describe, document, and register a Computational Workflow and a Computational Tool. The schema was presented using Schema.org, a web metadata markup standard, as part of the Bioschemas community, which produces subsets of Schema.org that are suitable for bioscience resources. The developed Bioschemas markup enables metadata to be more easily discovered by search engines and other aggregators such as Google and OpenAIRE.

While most life science services are openly accessible to anyone around the world, many of these require researchers to sign in using a username and password. Sensitive data and licensed resources, however, require strong security for access. In these cases, research services have implemented local access management solutions and issued their own usernames and passwords. As a result, researchers quickly become overloaded when trying to remember numerous login credentials.
The Life Science Login (LS Login) enables researchers to use their home organisation credentials or community, or even commercial identities (e.g. ORCID, LinkedIn), to sign in and access the data and services they need. It also allows service providers (both in academia and industry) to control and manage access rights of their users and create different access levels for research groups or international projects. In this way, this service provides authentication and authorisation for multiple other services within the life sciences.
For example, two different researchers in the same university may be working on different European or national research projects and may need access to completely different data or compute resources. LS Login ensures that each researcher can access the right resources by using their university credentials, while making sure they can’t see the other person’s data.
EOSC-Life created two vital systems to support data authentication, authorisation, and harmonisation processes: ARIA and LS Login.
The ARIA user access management system provides RIs, facilities and user communities in the field of structural biology and related fields with access to a collection of cloud services.

During the funding period, EOSC-Life issued two Open Calls and two Internal Calls that encouraged researchers to submit project proposals with innovative ideas for sharing data, tools, and workflows.
The response to these Calls was considerable, resulting in nineteen (19) projects being funded overall in the areas of the digital life sciences and cloud data deployment. These included projects that encouraged collaboration between academia and industry, proposed new ways to protect sensitive data, increased (the access to) the digital space for the life sciences, and proposed innovative methods to deploy data in the cloud.
Overview of these projects and their outcomes
This final report from WP3 provides a concise overview of the impacts.
Read about the Digital Life Sciences Open Call and funded projects
Three EOSC-Life Training Open Calls were issued from 2020-2023, and overall 11 scientific user projects were supported.
The Open Calls enabled EOSC-Life to achieve a practical goal: to support the co-creation of an open, digital, collaborative space for life science research by developing FAIR tools, workflows, resources, infrastructures, and guidelines with the help of the EOSC-Life experts and communities at the RIs.
Overviews of these projects and their impacts are provided on the EOSC-Life website at https://www.eosc-life.eu/services/open-call-training/ and in the final report from WP9 on EOSC-Life training activities.
Of particular interest were the 8 Demonstrator projects which provided the first concrete use cases in the initial phase of EOSC-Life.
EOSC-Life offered a variety of trainings that benefit both our project participants and the research community as a whole.
Links to training outcomes and resources are found at https://www.eosc-life.eu/services/training/.

All publications and deliverables produced by EOSC-Life can be found:
View all Project Deliverables & Publications